- All organisms shed traces of their biological material, which contains their unique DNA, into the environment. Researchers using a technique called metabarcoding to sequence this environmental DNA (eDNA) in water can detect the presence of multiple taxa in a single sample.
- eDNA surveillance is already being used as a tool for detecting invasive species and confirming the presence of endangered or cryptic organisms in an area, thereby influencing management decisions.
- Recent studies suggest that eDNA metabarcoding has the potential to support conservation efforts as a biodiversity monitoring tool in the marine environment.
- eDNA metabarcoding of water samples has proven to be an effective, non-invasive survey technique that allows researchers to assess the biodiversity of an aquatic environment in a fraction of the time that traditional manual survey methods require.
The Census of Marine Life (CoML), a 10-year international effort to determine the diversity of life found in Earth’s oceans, described over 1 million species, ranging from single-celled microbes to marine mammals the size of school buses. Understanding what species are present in our oceans, how abundant they are and how they are distributed is essential for developing effective conservation strategies. However, the vastness and complexity of the marine environment, coupled with this abundance of unique species, makes accurate and efficient monitoring of biodiversity a challenge.

The development of remote-sensing satellite technologies has revolutionized the collection of physiochemical ocean properties, such as temperature and wave-height. But, until recently, the collection of biological data has relied on labor-intensive survey methods that identify organisms based on morphological traits (shape, size, color etc.). Emerging technology based on the detection of environmental DNA (eDNA) may soon change that.
Environmental DNA (eDNA)
As organisms interact with their environment, they inevitably shed traces of biological material (skin, scales, feces, gametes, etc.) that contains their unique DNA. Since plant DNA was first detected in soil in 1998, eDNA from both plants and animals has been successfully isolated from samples collected in a variety of terrestrial and aquatic ecosystems. Traditionally, researchers have analyzed these samples using probes designed to detect the presence of a specific species in a process called DNA barcoding. This process is equivalent to that of a grocery store cashier who scans the barcode sticker on a costumer’s apple to determine its variety.
Marine researchers have used eDNA barcoding to confirm the presence of rare or cryptic organisms and to facilitate early detection of invasive species and harmful algal blooms. In addition, as highlighted in a recent Wildtech article, the Barcode of Wildlife Project (BWP) uses DNA barcoding to provide evidence in cases of poaching and illegal wildlife trade.

However, many important management and policy decisions depend on knowing the full suite of species present in a given area and their abundances. An emerging technology called eDNA metabarcoding may be able to provide that valuable information. Two recent studies have assessed the effectiveness of this biodiversity monitoring tool that has the potential to revolutionize how we are able to study the ocean and other ecosystems.
What is eDNA metabarcoding?
Although every individual organism contains a unique genetic code within their DNA, there are some areas of the DNA sequence, called “barcodes,” that are not only shared by individuals within the same species, but also across broader taxonomic groups. For example, the mitochondrial gene cytochrome c oxidase 1 (CO1) plays an essential role in energy production and is therefore highly conserved and found in almost all organisms.
In contrast to DNA barcoding, the process of metabarcoding is equivalent to the grocery cashier scanning and determining the variety of all of a customer’s produce, at the same time. Probes called primers are used to extract a specific barcode, such as CO1, from the DNA found in an environmental sample. A highly efficient process called high-throughput sequencing is able to rapidly determine the sequences of these barcodes and, by matching them to a reference database, researchers are able to determine which taxa are present.
The effectiveness of this technique lies in these special primers. While primers designed to sequence the CO1 barcode are highly effective at detecting eukaryotic DNA, more sensitive primers can be designed if researchers are interested in targeting a more specific taxonomic group with higher resolution. For example, in 2015 MiFish primers were developed for the universal detection of fish species. In a controlled aquarium experiment, eDNA metabarcoding using these primers successfully detected 168 out of 180 taxonomically diverse fish species.
eDNA metabarcoding as a biodiversity monitoring tool
A recent study in Scientific Reports compared the effectiveness of eDNA metabarcoding using MiFish primers to traditional underwater visual surveying. The researchers tested the new technique in the species-rich Maizuru Bay, in the Sea of Japan, where over 80 species of fish had been detected from 140 underwater visual surveys conducted over 14 years. Using eDNA metabarcoding in the same bay, researchers detected the presence of 112 species of fish from water samples. Satoshi Yamamoto and co-authors suggest that “this efficiency is potentially important, particularly in species-rich waters, because a greater effort is required to investigate the whole fish community as the number of species in the community increases.”
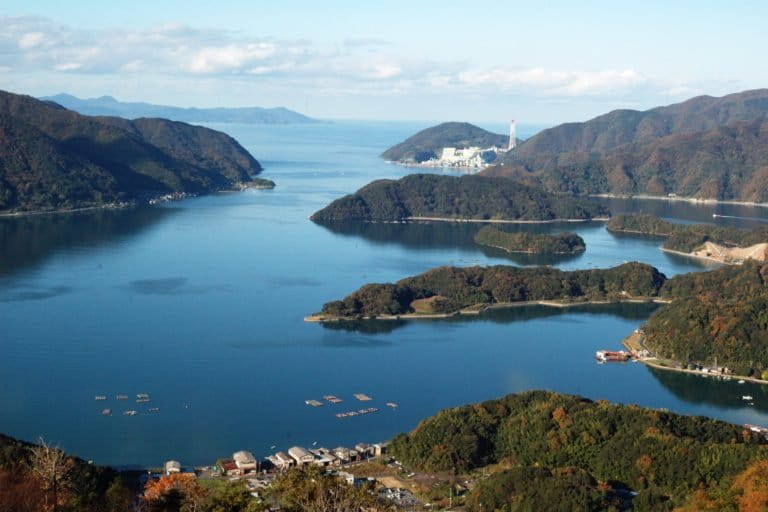

Not only was the eDNA metabarcoding technique more time-efficient and sensitive, but researchers detected the presence of 23 species that were not observed during the visual surveys. It is likely that some of these species were present in larval form – too small to be observed visually – or are well camouflaged. In addition, by sampling across a grid, the researchers were able to assess the utility of eDNA metabarcoding to estimate the spatial structure of the fish community. Most species were detected in their expected habitats, suggesting that the fish eDNA was not being transported far from its source organism.
A similar study recently published in Frontiers in Marine Science, also found that eDNA metabarcoding detected a broader range of animal taxa in seagrass habitats in Puget Sound, Washington, USA than did field surveys carried out using a manual-tow net. Using primers that targeted three different barcodes, University of Washington’s Ryan Kelly and colleagues sequenced eDNA from seawater samples and identified 366 taxonomic families, including various barnacles, snails, crabs, fish and seals. In contrast, the 12 manual tows conducted at the same locations identified just 45 distinct animal taxa (in 32 different families).
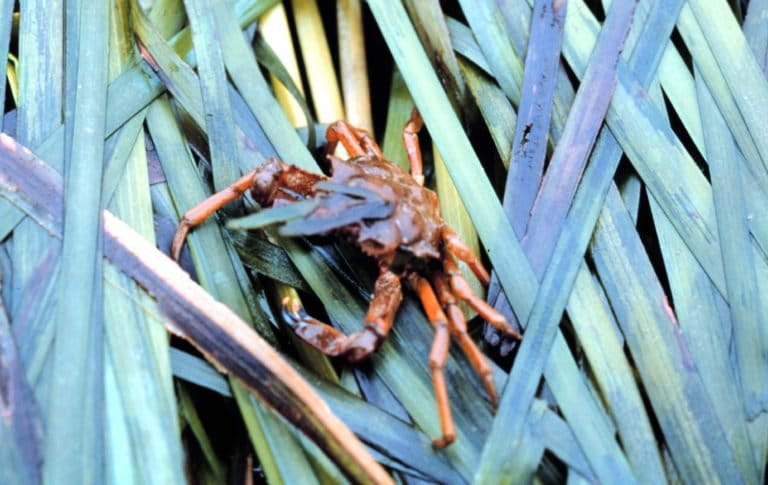

Analyses using the three different barcode regions and the manual tows also identified different taxa. “Our data suggest that different primer sets reveal different draws from a common pool of species represented in the sampled bottle of water,” the researchers concluded, adding.
They added, “These results highlight the value of using multiple methods in ecological surveys, given that any one sampling method—even eDNA, which can reveal hundreds of taxa present at a location— unavoidably reflects only a small fraction of the true biological diversity present in the environment.”

When feasible, they suggested, it helps to use complementary data types to assess biodiversity. “One advantage of using multiple data types is the ability to see deeper into an ecosystem than would otherwise be possible, each providing a new window into a complex living world.”
Pros and cons of eDNA metabarcoding
Metabarcoding of aquatic environmental DNA has proven to be a sensitive, non-invasive survey technique that allows researchers to assess the biodiversity of a given area in a fraction of the time that traditional survey methods require. Unlike terrestrial ecosystems, where eDNA has been shown to persist for years, aquatic ecosystems facilitate the rapid breakdown of eDNA. This reduces long-distance dispersal, giving researchers a more accurate snapshot of the species present and their spatial distribution.
In an e-mail to Wildtech, Kelly highlighted the pros of eDNA metabarcoding as “greatly increased sensitivity (ability to detect species when they are present) and coverage (number of species detected), as well as the ability to add more samples without greatly increasing cost.” Kelly also suggested, “There have now been dozens of validation studies in seawater, rivers, lakes, soil, air, etc. So it’s clear that this is a technique that is valuable and informative. But understanding the spatial and temporal resolution of eDNA surveys in each of these contexts is important, as is working to make the technique more quantitative.”
Monitoring an area’s biodiversity is essential for informing management and policy decisions; however, having some knowledge of species abundance is also often necessary. As Kelly suggests, compared to traditional survey methods, eDNA metabarcoding is still lacking in this quantitative aspect: “more DNA = more of a species, but we can’t say that 100 eDNA reads = 1 dolphin, for example,” he wrote.
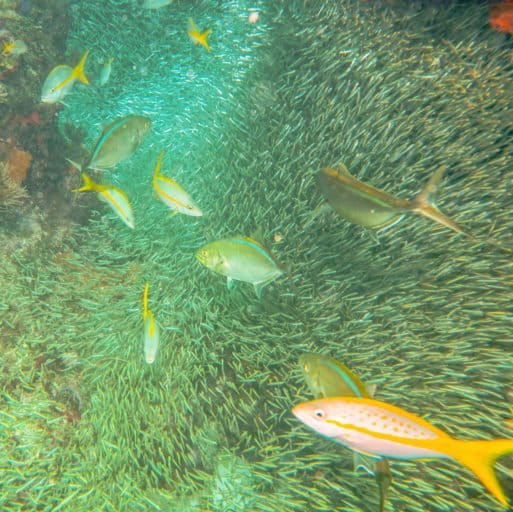

Kelly identified “the trade-off between how many species you’d like to detect and how much detail you would like to get about any particular species. So if you’re interested in the [genetic] diversity of a population of a single species, you can design eDNA probes that target that particular species (excluding all other species)…. Alternatively, if you want to survey a whole community (many species), you will lose information about diversity within populations of a particular species.”
Another potential drawback of eDNA barcoding highlighted by Kelly is that “it takes specialized training (not everyone is a molecular biologist with access to a lab).” Fortunately, increased interest in genetic research over the past decade has led to the development of technology that has significantly reduced the cost of sequencing. Ten years ago, it would have cost around $500 to obtain approximately 2000 DNA barcode sequences, as opposed to just $0.01 in 2017.
Reference databases (e.g. Genbank) used to match DNA sequences to species still use centuries of knowledge gained from visual identification. According to Kelly, “we very much rely on the natural history information — and the taxonomic expertise — of those ecologists that are in the field using traditional methods. I can’t imagine replacing that. But for applications where we can’t afford to pay people to do such labor-intensive sampling, some degree of automation makes sense, and eDNA might provide that.”

The future of eDNA metabarcoding
How long will it be until eDNA metabarcoding becomes an established and reliable method used to inform management and conservation policies? Perhaps not long. Environmental DNA surveillance has already proven to be a useful tool for detecting invasive and endangered species, thereby influencing management decisions. According to Kelly, “I think it’s just a matter of matching the questions that state/federal agencies are actually asking with the technology useful for answering those questions. Which is a process, but it’s already started. I get phone calls all the time from agencies asking how they can start using eDNA.”
Citations
Kelly RP, Closek CJ, O’Donnell JL, Kralj JE, Shelton AO and Samhouri JF (2017) Genetic and Manual Survey Methods Yield Different and Complementary Views of an Ecosystem. Frontiers in Marine Science. 3:283. doi: 10.3389/fmars.2016.00283
Yamamoto, S. et al. (2017) Environmental DNA metabarcoding reveals local fish communities in a species-rich coastal sea. Scientific Reports. 7:40368. doi: 10.1038/srep40368







