- The Global Biodiversity Information Facility (GBIF) is now the largest biodiversity database in the world with records of hundreds of millions of occurrences of over 1.7 million species, ranging from bacteria to blue whales.
- Institutions from over 50 countries contribute species occurrence and related data to the open-access platform, which make possible regional or global-scale analyses of data ranging from global distributions to invasive species and climate change impacts.
- As GBIF and other collaborative, open-source data bases continue to expand and mature, so will their usefulness to a greater range of scientific studies.
How many species are living on Earth at this moment in time? Ask a few different scientists and you may get drastically different answers. Most estimates range from 3-10 million distinct species of multicellular organisms; however, when microbial diversity is factored in, the upper bound jumps to nearly a trillion.
Having a clear understanding of what species are present on Earth, where they are, and to what extent they are threatened is essential to making informed conservation decisions at both a local and global scale.
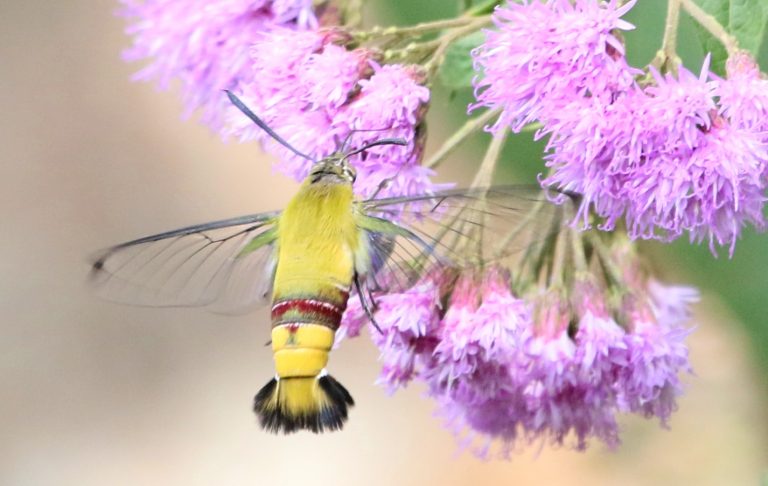

In 2001, the Global Biodiversity Information Facility (GBIF) was established through the signing of a Memorandum of Understanding by participating countries with the goal of providing the infrastructure to store biodiversity datasets in a standardized format that is accessible to everyone. The overall vision of GBIF is to have “A world in which biodiversity information is freely and universally available for science, society and a sustainable future”.
GBIF has now grown into the largest biodiversity database in the world with records of hundreds of millions of occurrences of over 1.7 million species, ranging from bacteria to blue whales. It contains data collected over the past three centuries from biological surveys as well as digitized records of museum and herbarium collections. Institutions from over 50 countries have contributed datasets to GBIF.

How it works
Participating countries are encouraged to create a ‘node’ to help coordinate the work of GBIF in that country. This node can be part of an existing institution that already manages biodiversity data, or an independent entity. These nodes help to coordinate the upload of that country’s biodiversity to the GBIF in a standardized format.
Uploading data requires an endorsement from GBIF that ensures the data are relevant and available for open access. Citizen scientists can upload their observation data through established networks such as the ArtDatabanken in Sweden or through mobile applications such as eBird and iNaturalist. In keeping with the GBIF’s mission, all users uploading data to the GBIF must agree to make their contributions open to reuse by others.
The GBIF supports the publishing of four classes of datasets – resource metadata, checklist data, occurrence-only data and sampling-event data. The bulk of the data in the GBIF comprise species occurrence observations that document when and where a species was found with specificity ranging from a GPS point to a country. Sampling-event datasets, obtained through standardized protocols such as vegetation transects, document community composition at different locations and/or times.
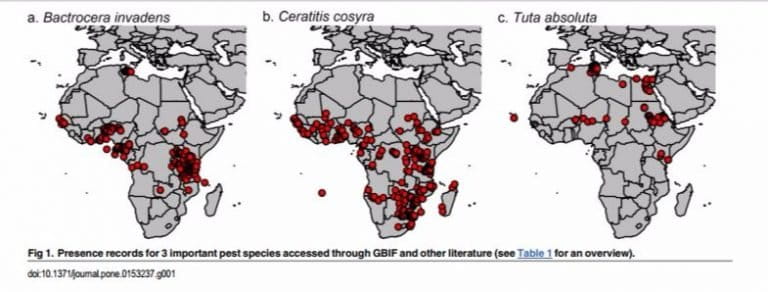
Users can access data through GBIF.org with the option of searching based on taxon or country. When occurrence data are georeferenced with GPS coordinates, users can map species distributions directly on the GBIF website using optional location and time filters.
Users can download data as a simple CSV file, which gives a tabular view of the data, or as a Darwin Core Archive zip file, which includes additional information such as images. Researchers who download data from the GBIF must follow the data user agreement which ensures that proper credit is given to the publisher of the dataset.
Moving biodiversity research forward
Having free and unlimited access to the extensive GBIF biodiversity database has allowed scientists to ask research questions that previously would have been impossible to investigate due to data constraints. To date, over 1,400 peer-reviewed research publications have cited the GBIF as a data source and, on average, about one paper citing GBIF is now published each day.
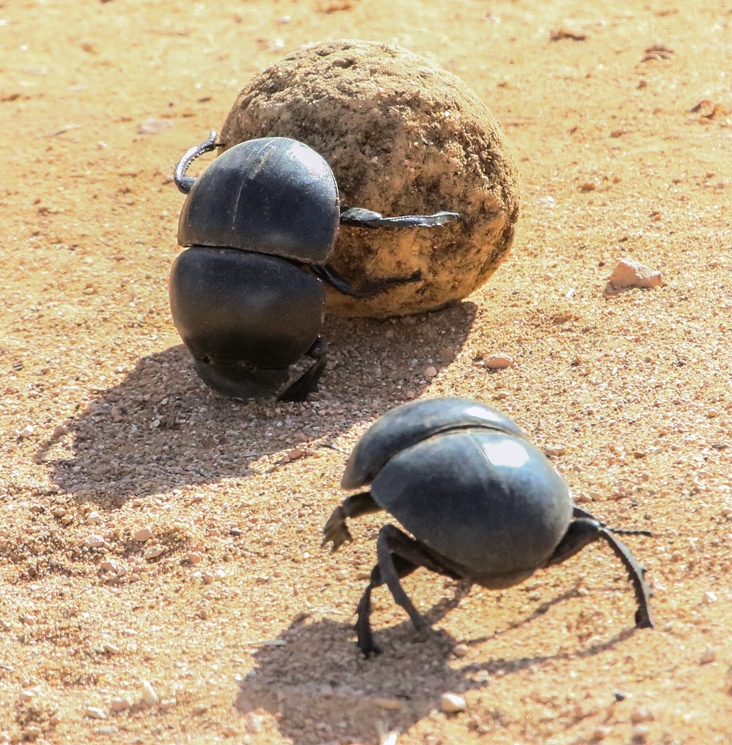

Dr. Celine Bellard, a researcher at the Centre for Biodiversity & Environment Research at the University College of London, uses GBIF species occurrence datasets to determine the influence of environmental factors on invasive species distributions.
In an email to Mongabay-Wildtech, Bellard stated, “Without the GBIF, I could not conduct the study that I am doing. Because it would not be possible to collect all those data by myself and then work on such project.”
Other researchers have used the open database for a variety of different applications, which are highlighted in the GBIF’s annual science review. Species occurrence data in the form of GPS coordinates are used by researchers across the globe to map species distributions. Occurrence data can also be used to perform biodiversity assessments to identify priority areas for conservation. Researchers are also able to integrate environmental data with species occurrence data from GBIF in order to gain insight into the influence of land-use and climate change on species distributions.
Susan Canavan, a PhD candidate with the Centre for Invasion Biology, Department of Botany and Zoology at Stellenbosch University, used the GBIF in a recent study on the global distribution of bamboos. In an e-mail to Mongabay-Wildtech, Canavan stated, “Establishing up-to-date inventories of taxa and their distribution is fundamental for invasion science. Global biodiversity databases are therefore a valuable tool for helping us understand how humans have shuffled species around the world, and what the consequences are of this.”

GBIF data has also been used to study the risk of emergence of human zoonotic diseases. According to Dr. David Redding of the Center for Biodiversity & Environment Research at the University College of London, “GBIF is an unparalleled resource for the work I do. It effectively enables me to do my work – determining on a large geographic scale where disease-carrying, non-human species are likely located, to better understand where human disease burden is highest.”
In an e-mail to Mongabay-Wildtech, Dr. Biber-Freudenberger of the University of Bonn, who uses GBIF data to study the influence of climate change on species distributions, stated, “The compilation of existing data in large databases such as GBIF is the only way for researchers all over the world to access existing data. Efforts should be intensified to complete these databases as far as possible and feed all existing information into them.”
Areas for improvement
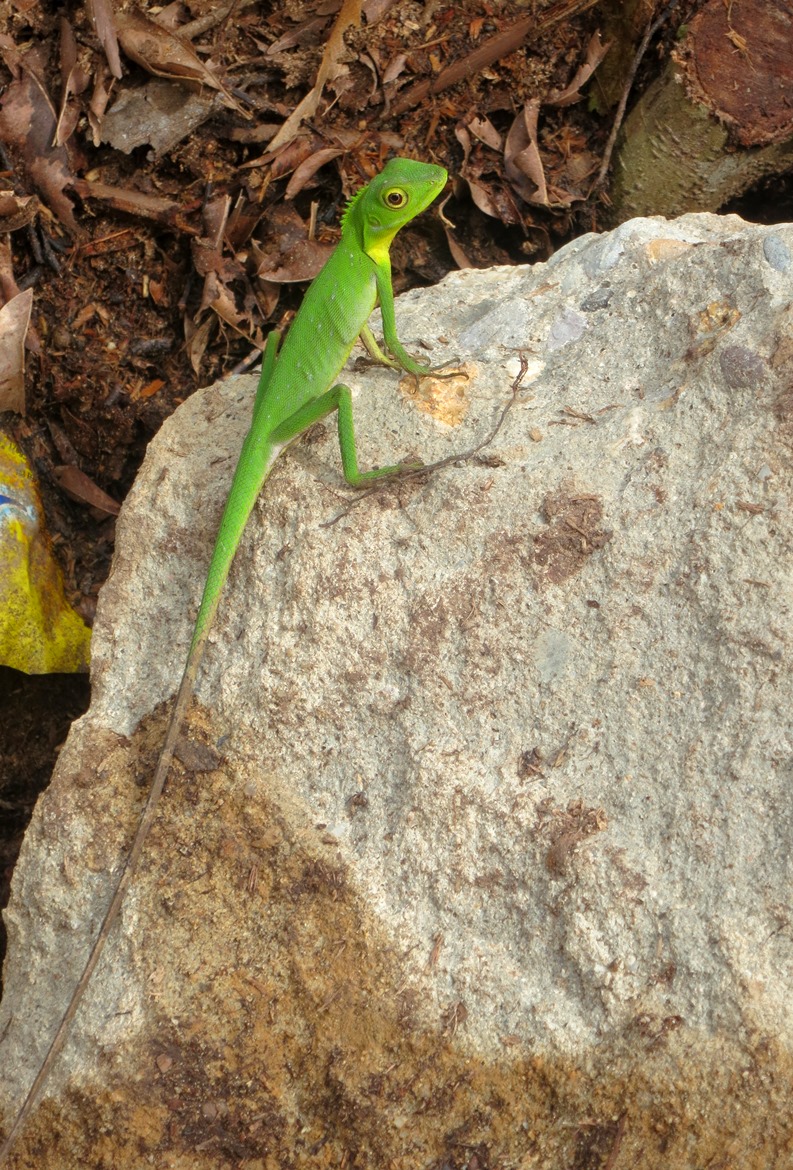

There are still areas where the database can be improved. According to Bellard, one helpful addition would be “to include information about the certainty of the data. Specifically regarding invasive species, information about the status at each location (invasive, established, failed, native) would be particularly useful.”
Redding added, “I would really like to know the precise environmental conditions where specimens were found i.e. rice paddy, primary woodland, house. I have to rely on coarse satellite data to guess at the best answer at the moment.”
“There could also be more precise spatial data for wild organisms, better quantification of the data biases, and more ways to interact with the temporal aspects of the data set. It would be great to have tools, for instance, that easily extract from environmental data both the location and the date.”
With the large number of datasets being uploaded each day, it is inevitable that some contain errors or incorrect formatting. According to Canavan, “One of the challenges of working with a database compiled from many different sources like GBIF is that there is often a large amount of cleaning that is required to improve the quality of the data. Incorrect GPS coordinates or synonyms are common issues. When you are interested in reviewing large sets of species, such as bamboos (c. 1600 species) that have over 100,000 records, cleaning can be a big task. Although, recently there have been efforts in developing ways to automate this process.”

Dr. Bradley Butterfield from the Department of Biological Sciences at Northern Arizona University told Mongabay-Wildtech how users are accessing large numbers of data records more efficiently.
He explained, “The standard HTML interface for GBIF is fine, particularly for exploring data, looking at geographic distributions of a single species, etc., but is not very useful for downloading records for many species simultaneously. However, there are a number of folks who have written code on various software platforms that make this really easy. For example, I use an R package called Dismo to quickly download thousands of records each for dozens to hundreds of species for various projects. So what I see as a primary limitation of GBIF has been solved by some rather clever people using open-source software.”
The future of the GBIF and global biodiversity research
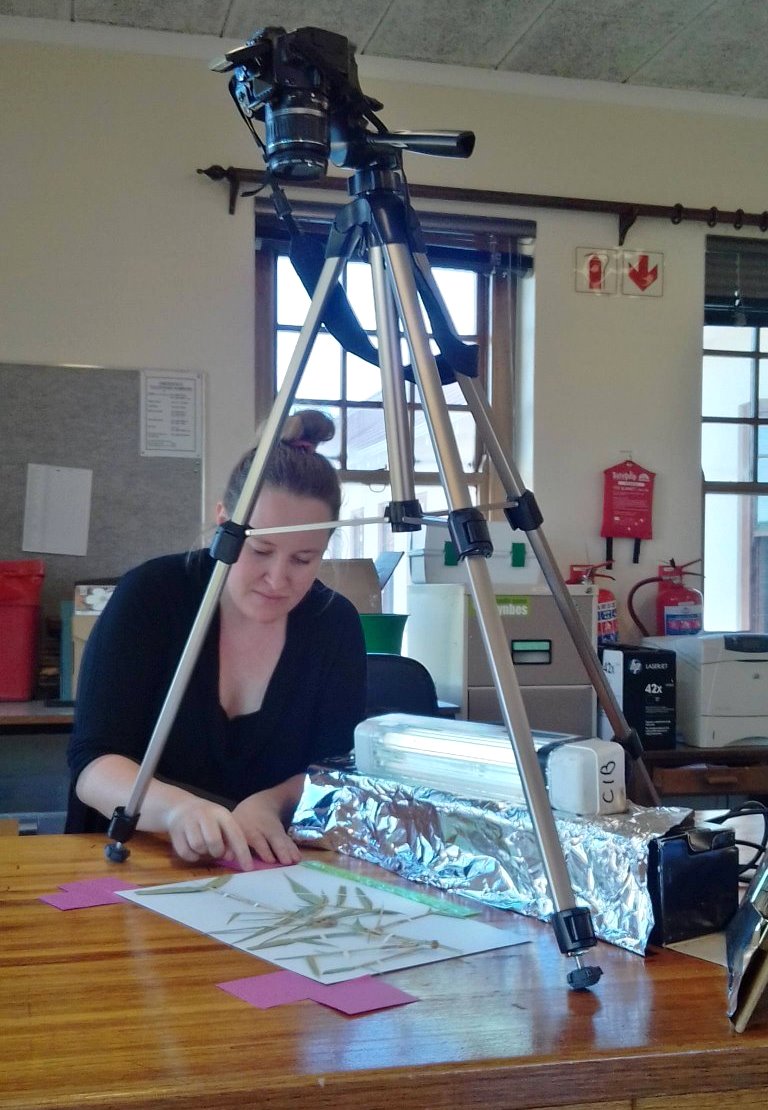

The GBIF has achieved its initial goal of creating an infrastructure for making biodiversity data accessible and usable for anyone. Looking forward, it is likely that open-source software like Dismo will help improve the usefulness of this integrated global dataset.
As highlighted in a previous Wildtech article, photographs taken by citizen scientists and tourists can become a substantial source of data for scientific studies when coupled with image identification software.
Redding also brought up photographs as a potential source of data for the GBIF, “Adding in sightings from social media, e.g. photos of zebra automatically identified using machine learning, would be a great way to start populating the GBIF database without the need for a large amount of human resources.”
As the GBIF continues to grow, so will its usefulness for a greater range of scientific studies. As expressed by Butterfield, “the ease with which data can be acquired, assessed and modeled via sources like GBIF is a great example of what collaborative science does and should look like in the 21st century.”

