- The recent DNA Barcode Scanner Hack brought together a range of experts to brainstorm a handheld modular DNA analysis device that could identify timber samples in the field, help flag wildlife trafficking, detect novel pathogens and enhance fisheries traceability, bypassing the need for an expensive, distant laboratory.
- Survey responses to the Wildtech needs assessment put portable DNA analysis near the top of the research and conservation technology wish-list.
- Future research and development will need to test all of the extraction methods for all of the main target species; this will require long-term vision, funding and central coordination across laboratories worldwide.
Survey responses to the Wildtech needs assessment—especially from field researchers—put portable DNA analysis near the top of the research and conservation technology wish-list. How could we harness DNA barcode technology in an adaptable, handheld device that frontline officials in developing countries could use to combat timber and wildlife trafficking?
The DNA Barcode Scanner Hack that took place in Washington, DC on Friday, August 29, dug into this question. The meeting brought together over 40 experts in evolution and molecular biology, conservation, design, engineering and enforcement to brainstorm an open-source technology that could identify timber samples in the field, help flag wildlife trafficking, detect novel pathogens and enhance fisheries traceability, bypassing the need for an expensive, distant laboratory. Conservation X Labs, Smithsonian Institution and the Consortium for the Barcode of Life (CBOL) hosted the event at the National Museum of American History.
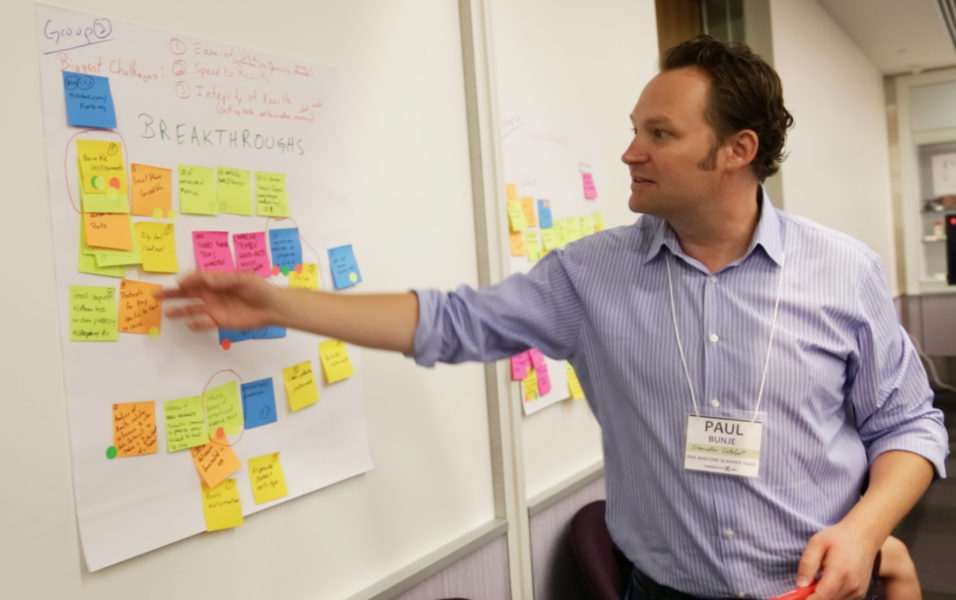

“The event was extraordinary,” said David Baisch, Molecular Innovations Fellow at Conservation X Labs. “We hoped that this event would identify clear development pathways for approaching a prototype which combined synergistic DNA isolation and analysis methods for timber, while keeping our design criteria of being cheap (a device cost of less than $1000, which could analyze a sample for $5 or less), portable, manufacturable, scalable, usable by an individual with no scientific training, in conditions without regular access to power, challenging environmental conditions, and limited distribution networks for reagents.”
Four challenges pertaining to developing a portable DNA reader structured the event: design criteria limitations and solutions, innovative timber DNA isolation, field-ready timber DNA sequence amplification and analysis technologies and, finally, solution integration and prototyping. These challenges aimed to help hackers identify the most effective means of creating the minimum viable prototype.
“These conversations were extremely productive, and this multifaceted approach to conservation technology design produced a large amount [number] of great ideas and approaches,” reflected Baisch, also noting that the meeting could have been week-long and given more time to design and testing.

As one of the conference’s attendees, Environmental Investigation Agency (EIA) Eurasia Programs Coordinator David Gehl brought to the table a background in tracing supply chains and pushing for policy that improves transparency.
“It was an excellent convening of some of the most relevant actors in a very structured way focused on developing innovative solutions to this critical problem,” he said.
According to Gehl, the meeting succeeded in creating a framework of concrete steps for tackling the complications surrounding the extraction of usable wood DNA in a handheld field-ready device.

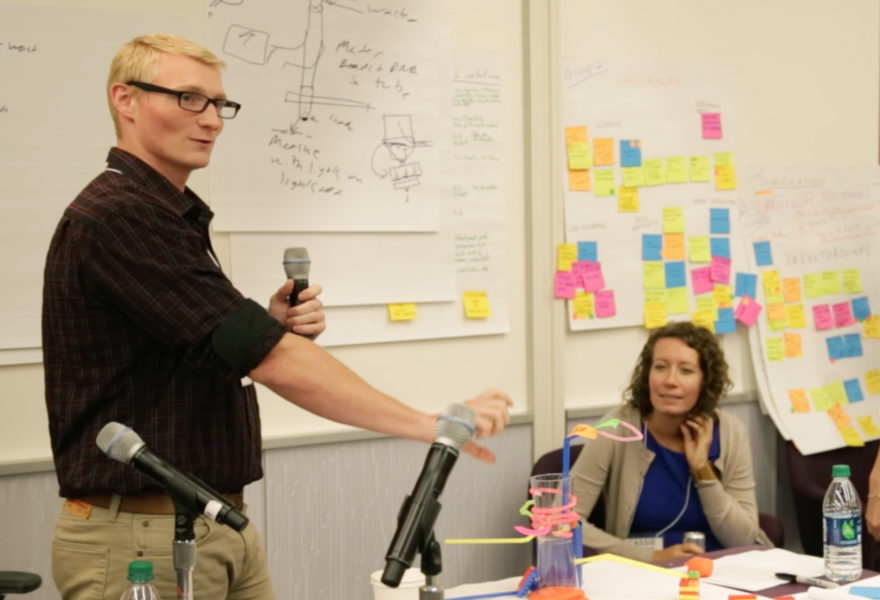
“The DNA barcode is a short gene sequence taken from a standard position in the genome,” explained David Schindel, Director of CBOL at the Smithsonian. “The region is selected as the barcode region because it has the closest concordance to the clusters we call species. The discontinuity between species corresponds to the discontinuity between DNA sequences.”
The envisioned DNA barcode scanner will utilize open reference databases from the University of Guelph’s Barcode of Life library, which have been completed for over 500,000 species. The library is based on the C01 gene, which has a unique sequence for every species, especially vertebrates, and therefore serves well as a barcode; however, datasets based on other sequences need to be developed for timber.
Conservation X Labs and partners are coming up with new techniques and turning to cutting-edge technologies to make the preparation and analysis of biological samples as simple as possible while weighing simplification against its potential ramifications for accuracy.
“These functionality requirements are challenging, but are absolutely surmountable,” said CEO of Conservation X Labs Alex Dehgan.
This conference focused on tree species identification to fight the illegal timber trade, but the technology could be slightly altered for applications in wildlife trafficking, disease surveillance, food safety and fish identification and traceability.

Possible alternatives
The market does include other powerful handheld devices, such as nanopore sequencers; however, these are too expensive to be feasible on the frontlines and haven’t been fully tested with plant tissue and its relevant challenges.
Still, avenues other than DNA barcoding, such as stable isotopes, near-infrared spectroscopy, mass spectrometry, wood anatomy and machine vision also hold promise. As Gehl put it, each technology has advantages over the others, depending on the particular question at hand.
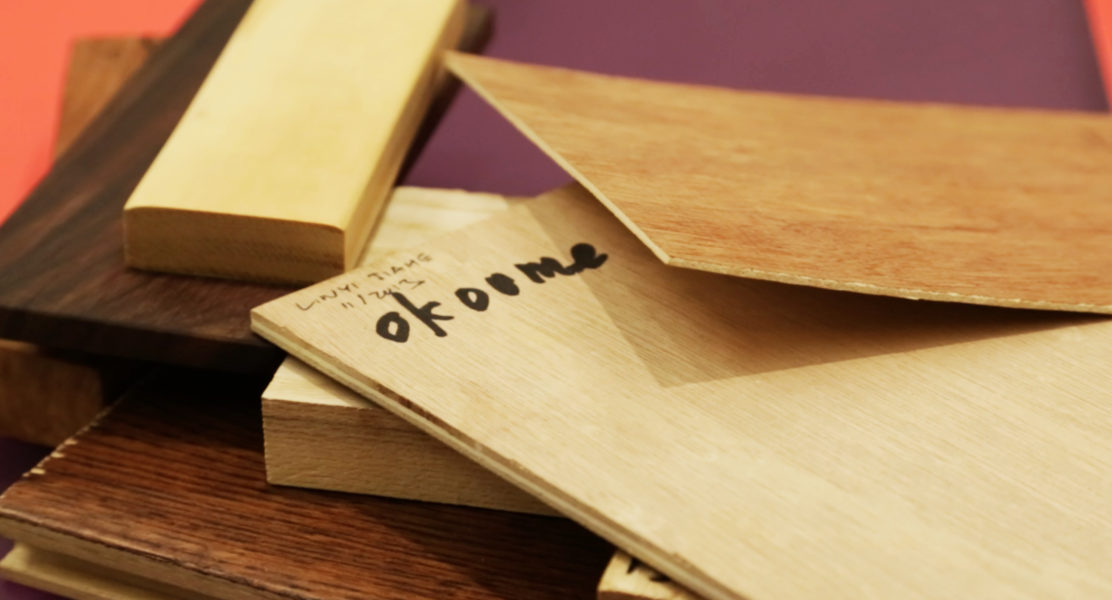
“The biggest hope is to develop a field-ready handheld device that can identify the species of wood in a piece of furniture or flooring coming in at the US border and to check that identification against the declared species and country of harvest listed on the Lacey Act declaration form,” he said. “Such a device can also be invaluable to help companies test products from suppliers, to make sure they’re getting what they paid for.”
Planning the conservation tech hack meeting
The Conservation X Labs DNA Barcode Scanner project began a few months ago. The hack event took just over a month of planning and was funded by a grant from the National Fish and Wildlife Foundation. The money came from Lumber Liquidators’ high-profile indictment last year for the company’s unlawful importation of hardwood illegally logged in the Russian Far East. The funds also cover research toward the development of timber identification technologies.
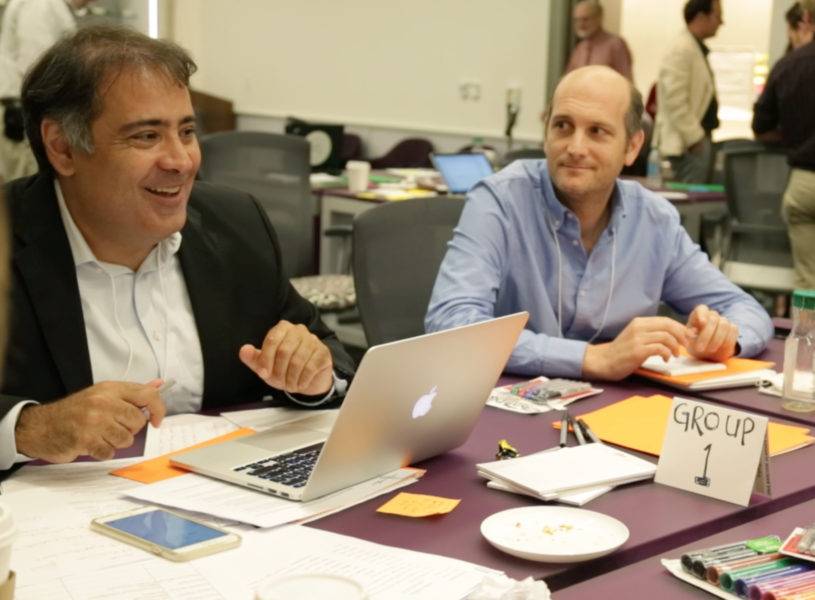

According to Dehgan, it had two main inspirations. One was University of Pennsylvania tropical biologist Dan Janzen’s vision of a tool that could use the Barcode of Life library to empower everyone from customs officials to children to better understand the living world. The organizers have been awaiting the development of such a DNA barcode scanner since hearing him put forward the idea at an American Association for the Advancement of Science Fellowship event a decade ago. Tired of waiting, they eventually decided to capitalize on the democratization of genomic technology and propel the scanner’s realization themselves.
The second impetus was the illegal timber and wildlife trades. The Lumber Liquidator’s case, for instance, illustrated the need for a way to help verify supply chains, conserve endangered species and enforce wildlife trade laws.
What came out of the conference?
“As with many groundbreaking scientific endeavors, we left the event with many more questions than we came into it with, but most of our initial questions were answered outright or now have answers on clear research pathways,” said Baisch.
“I came away from the event feeling more optimistic about the process. There is a path forward; there is interest,” added Gehl. “The technology is there; the ability to do the science and build the device exists. What will be critical is having the long-term vision and funding to establish a coordinated effort between different scientific laboratories.”
Gehl explained that the hack left the experts with a broader understanding that the project would have to encompass all of the extraction methods for all of the targeted main species and systematically test these methods. Because large-scale testing can be tedious and costly depending on the reagents used, it would be most efficient if conducted by many laboratories worldwide. As different methods work best for different species, products and DNA types, researchers must coordinate their efforts to avoid duplication and determine the most effective technique for each case. Such coordination requires help from Conservation X Labs or another external party capable of collating and analyzing results in a scientifically sound manner.
“What this project does is that essential step of looking at it all comprehensively and saying, what are the key issues to be resolved to make connections between labs and push research in the direction it needs to go to make this a successful technology?” Gehl said.

Conservation X Labs will keep convening the open source community to determine how to overcome constraints. The organization plans to hold a series of small challenges and work with some of the participants around various competing technologies.
“The real excitement rests in portable whole genome sequencing technology that can not only compare DNA samples against a known set of species, but conduct true biological discovery in the field,” concluded Dehgan. “This can unlock new drugs, insights into the biological world, serve as early detection of invasive species and emerging infectious diseases, and help us protect and sustain this planet’s biodiversity, including that of our own species.”







